|
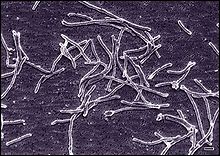
Thermus aquaticus
Thermus aquaticus is a species of bacteria that can tolerate high temperatures, one of several thermophilic bacteria that belong to the Deinococcota phylum. It is the source of the heat-resistant enzyme Taq DNA polymerase, one of the most important enzymes in molecular biology because of its use in the polymerase chain reaction (PCR) DNA amplification technique. History When studies of biological organisms in hot springs began in the 1960s, scientists thought that the life of thermophilic bacteria could not be sustained in temperatures above about 55 °C (131 °F).[1] Soon, however, it was discovered that many bacteria in different springs not only survived, but also thrived in higher temperatures. In 1969, Thomas D. Brock and Hudson Freeze of Indiana University reported a new species of thermophilic bacteria which they named Thermus aquaticus.[2] The bacterium was first isolated from Mushroom Spring in the Lower Geyser Basin of Yellowstone National Park, which is near the major Great Fountain Geyser and White Dome Geyser,[3] and has since been found in similar thermal habitats around the world. Decades later, this discovery had profound implications, including the invention of Polymerase Chain Reaction (PCR) by biochemist Kary Mullis, which revolutionized DNA research and earned Mullis a Nobel Prize in Chemistry in 1993. PCR facilitated advancements in medical diagnostics, genetics, and other fields. After Kary Mullis' discovery of PCR, Cetus awarded him $10,000. However, Cetus later sold the PCR patent to F. Hoffmann-La Roche (Roche) for $300 million. This transaction left Mullis feeling cheated throughout his life. Roche has since profited immensely from the PCR patent, with annual PCR-related sales reaching $5.4 billion in 2022. Despite these profits, neither the National Park Service, Yellowstone National Park, nor the state of Wyoming have received any share of these revenues. Recognizing the scientific and financial potential of Yellowstone's extremophiles, biotechnology companies like Diversa signed agreements with the National Park Service for bioprospecting. This led to further scientific exploration and potential commercial applications, despite some environmental concerns. Overall, Brock's initial discovery in Yellowstone's hot springs paved the way for significant scientific breakthroughs, demonstrating the importance of basic research in driving innovation and technological advancements.[4] BiologyT. aquaticus shows best growth at 65–70 °C (149–158 °F), but can survive at temperatures of 50–80 °C (122–176 °F). It primarily scavenges for protein from its environment as is evidenced by the large number of extracellular and intracellular proteases and peptidases as well as transport proteins for amino acids and oligopeptides across its cell membrane. This bacterium is a chemotroph—it performs chemosynthesis to obtain food. However, since its range of temperature overlaps somewhat with that of the photosynthetic cyanobacteria that share its ideal environment, it is sometimes found living jointly with its neighbors, obtaining energy for growth from their photosynthesis. T. aquaticus normally respires aerobically but one of its strains, Thermus aquaticus Y51MC23, is able to be grown anaerobically.[5] The genetic material of T. aquaticus consists of one chromosome and four plasmids, and its complete genome sequencing revealed CRISPR genes at numerous loci.[6] MorphologyThermus aquaticus is generally of cylindrical shape with a diameter of 0.5 μm to 0.8 μm. The shorter rod shape has a length of 5 μm to 10 μm. The longer filament shape has a length that varies greatly and in some cases exceeds 200 μm. T. aquaticus has shown multiple possible morphologies in different cultures, rod-shaped or as short filaments. The rod-shaped bacteria have a tendency to aggregate. Associations of several individuals can lead to the formation of spherical bodies 10 μm to 20 μm in diameter, also called rotund bodies.[2][7] These bodies are not composed of cell envelope or outer membrane components as previously thought, but are instead made from remodelled peptidoglycan cell wall. Their exact function in the survival of T. aquaticus remains unknown but has been theorised to include temporary food and nucleotide storage, or they may play a role in the attachment and organisation of colonies.[6] Thermus aquaticus is a typical gram-negative bacterium, which indicates that its cell walls have considerably less peptidoglycan compared to gram-positive counterparts. In the presence of sunlight, Thermus can display hues ranging from yellow to pink or red, which are visible in hot springs. Additionally, Thermus aquaticus may possess flagella for motility or remain immotile. Enzymes from T. aquaticusT. aquaticus has become famous as a source of thermostable enzymes, particularly the Taq DNA polymerase, as described below. AldolaseStudies of this extreme thermophilic bacterium that could be grown in cell culture was initially centered on attempts to understand how enzymes, which are normally inactive at high temperature, can function at high temperature in thermophiles. In 1970, Freeze and Brock published an article describing a thermostable aldolase enzyme from T. aquaticus.[8] RNA polymeraseThe first polymerase enzyme isolated from T. aquaticus in 1974 was a DNA-dependent RNA polymerase,[9] used in the process of transcription. Taq I restriction enzymeMost molecular biologists probably became aware of T. aquaticus in the late 1970s or early 1980s because of the isolation of useful restriction endonucleases from this organism.[10] Use of the term Taq to refer to Thermus aquaticus arose at this time from the convention of giving restriction enzymes short names, such as Sal and Hin, derived from the genus and species of the source organisms. DNA polymerase ("Taq pol")DNA polymerase was first isolated from T. aquaticus in 1976.[11] The first advantage found for this thermostable (temperature optimum 72°C, does not denature even in 95 °C) DNA polymerase was that it could be isolated in a purer form (free of other enzyme contaminants) than could the DNA polymerase from other sources. Later, Kary Mullis and other investigators at Cetus Corporation discovered this enzyme could be used in the polymerase chain reaction (PCR) process for amplifying short segments of DNA,[12] eliminating the need to add E. coli polymerase enzymes after every cycle of thermal denaturation of the DNA. The enzyme was also cloned, sequenced, modified (to produce the shorter 'Stoffel fragment'), and produced in large quantities for commercial sale.[13] In 1989 Science magazine named Taq polymerase as its first "Molecule of the Year".[14] In 1993, Mullis was awarded the Nobel Prize in Chemistry for his work with PCR.[15] Other enzymesThe high optimum temperature for T. aquaticus allows researchers to study reactions under conditions for which other enzymes lose activity. Other enzymes isolated from this organism include DNA ligase, alkaline phosphatase, NADH oxidase, isocitrate dehydrogenase, amylomaltase, and fructose 1,6-disphosphate-dependent L-lactate dehydrogenase. ControversyThe commercial use of enzymes from T. aquaticus has not been without controversy. After Brock's studies, samples of the organism were deposited in the American Type Culture Collection, a public repository. Other scientists, including those at Cetus, obtained it from there. As the commercial potential of Taq polymerase became apparent in the 1990s,[16] the National Park Service labeled its use as the "Great Taq Rip-off".[17] Researchers working in National Parks are now required to sign "benefits sharing" agreements that would send a portion of later profits back to the Park Service. See alsoReferences
Further reading
External links |
||||||||||||||||||||||||||||