|
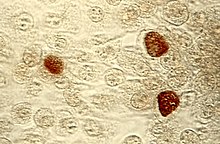
Chlamydia trachomatis
Chlamydia trachomatis (/kləˈmɪdiə trəˈkoʊmətɪs/) is a Gram-negative, anaerobic bacterium responsible for chlamydia and trachoma. C. trachomatis exists in two forms, an extracellular infectious elementary body (EB) and an intracellular non-infectious reticulate body (RB).[2] The EB attaches to host cells and enter the cell using effector proteins, where it transforms into the metabolically active RB. Inside the cell, RBs rapidly replicate before transitioning back to EBs, which are then released to infect new host cells.[3] The earliest description of C. trachomatis was in 1907 by Stanislaus von Prowazek and Ludwig Halberstädter as a protozoan.[4] It was later thought to be a virus due to its small size and inability to grow in laboratories. It was not until 1966 when it was discovered as a bacterium by electron microscopy where its internal structures were observed. There are currently 18 serovars of C. trachomatis, each associated with specific diseases affecting mucosal cells in the lungs, genital tracts, and ocular systems.[3] Infections are often asymptomatic, but can lead to severe complications such as pelvic inflammatory disease in women and epididymitis in men. The bacterium also causes urethritis, conjunctivitis, and lymphogranuloma venereum in both sexes. C. trachomatis genitourinary infections are diagnosed more frequently in women than in men, with the highest prevalence occurring in females aged 15 to 19 years of age.[5][6][7] Infants born from mothers with active chlamydia infections have a pulmonary infection rate that is less than 10%.[8] Globally, approximately 84 million people are affected by C. trachomatis eye infections, with 8 million cases resulting in blindness.[9] C. trachomatis is the leading infectious cause of blindness and the most common sexually transmitted bacterium.[3] The impact of C. trachomatis on human health has been driving vaccine research since its discovery.[10] Currently, no vaccines are available, largely due to the complexity of the immunological pathways involved in C. trachomatis, which remain poorly understood. However, C. trachomatis infections may be treated with several antibiotics, with tetracycline being the preferred option.[11][12] DescriptionChlamydia trachomatis is a gram-negative bacterium that can replicate exclusively within a host cell, making it an obligate intracellular pathogen.[3] Over the course of its life cycle, C. trachomatis takes on two distinct forms to facilitate infection and replication. Elementary bodies (EBs) are 200 to 400 nanometers across and are surrounded by a rigid cell wall that enables them to survive in an extracellular form.[3][10] When an EB encounters a susceptible host cell, it binds to the cell surface and is internalized.[3] The second form, reticulate bodies (RBs) are 600 to 1500 nanometers across, and are found only within host cells.[10] RBs have increased metabolic activity and are adapted for replication. Neither form is motile.[10] The evolution of C. trachomatis includes a reduced genome of approximately 1.04 megabases, encoding approximately 900 genes.[3] In addition to the chromosome that contains most of the genome, nearly all C. trachomatis strains carry a 7.5 kilobase plasmid that contains 8 genes.[10] The role of this plasmid is unknown, although strains without the plasmid have been isolated, suggesting it is not essential for bacterial survival.[10] Several important metabolic functions are not encoded in the C. trachomatis genome and are instead scavenged from the host cell.[3] Carbohydrate metabolismC. trachomatis has a reduced metabolic capacity due to its smaller genome, which lacks genes for many biosynthetic pathways including those required for complete carbohydrate metabolism.[3][13] The bacterium is largely dependent on the host cell for metabolic intermediates and energy, particularly in the form of adenosine triphosphate (ATP).[13] C. trachomatis lacks several enzymes necessary for independent glucose metabolism, and instead utilizes two ATP/ADP translocases (Npt1 and Npt2) to import ATP from the host cell.[14] Other metabolites including amino acids, nucleotides, and lipids are also transported from the host.[13][15] A critical enzyme involved in glycolysis, hexokinase, is absent in C. trachomatis, preventing the production of glucose-6-phosphate (G6P). Instead, G6P from the host cell is taken up by the metabolically active reticulate bodies (RBs) through a G6P transporter (UhpC antiporter).[15][13] Although C. trachomatis lacks a complete independent glycolysis pathway, it has genes encoding for all the enzymes required for the Pentose Phosphate Pathway (PPP), gluconeogenesis, and glycogen synthesis and degradation.[13] A suppressor of glycolysis, p53, is expressed less frequently in c. trachomatis cells, increasing the rate at which glycolysis occurs, even in the presence of oxygen.[13] As a result, C. trachomatis infection is associated with increased production of pyruvate, lactate, and glutamate by the bacterium due to activity of the pyruvate dehydrogenase kinase 2 enzyme limiting conversion of pyruvate to acetyl-coenzyme A.[13] The pyruvate is instead turned into lactate, which allows the cell to grow almost unobstructed by immune response due to the acidic properties of lactate.[13][16] Excess glycolytic products are, in turn, brought into the PPP to create nucleotides and for biosynthesis.[13] This type of growth is very similar to the Warburg effect observed in cancer cells.[13] Life cycle Like other Chlamydia species, C. trachomatis has a life cycle consisting of two morphologically distinct forms. First, C. trachomatis attaches to a new host cell as a small spore-like form called the elementary body.[17] The elementary body enters the host cell, surrounded by a host vacuole, called an inclusion.[17] Within the inclusion, C. trachomatis transforms into a larger, more metabolically active form called the reticulate body.[17] The reticulate body substantially modifies the inclusion, making it a more hospitable environment for rapid replication of the bacteria, which occurs over the following 30 to 72 hours.[17] The massive number of intracellular bacteria then transition back to resistant elementary bodies before causing the cell to rupture and being released into the environment.[17] These new elementary bodies are then shed in the semen or released from epithelial cells of the female genital tract and attach to new host cells.[18] ClassificationC. trachomatis are bacteria in the genus Chlamydia, a group of obligate intracellular parasites of eukaryotic cells.[3] Chlamydial cells cannot carry out energy metabolism and they lack biosynthetic pathways.[19] C. trachomatis strains are generally divided into three biovars based on the type of disease they cause. These are further subdivided into several serovars based on the surface antigens recognized by the immune system.[3] Serovars A through C cause trachoma, which is the world's leading cause of preventable infectious blindness.[20] Serovars D through K infect the genital tract, causing pelvic inflammatory disease, ectopic pregnancies, and infertility. Serovars L1 through L3 cause an invasive infection of the lymph nodes near the genitals, called lymphogranuloma venereum.[3] C. trachomatis is thought to have diverged from other Chlamydia species around 6 million years ago. This genus contains a total of nine species: C. trachomatis, C. muridarum, C. pneumoniae, C. pecorum, C. suis, C. abortus, C. felis, C. caviae, and C. psittaci. The closest relative to C. trachomatis is C. muridarum, which infects mice.[17] C. trachomatis along with C. pneumoniae have been found to infect humans to a greater extent. C. trachomatis exclusively infects humans. C. pneumoniae is found to also infect horses, marsupials, and frogs. Some of the other species can have a considerable impact on human health due to their known zoonotic transmission.[3]
Role in diseaseClinical signs and symptoms of C. trachomatis infection in the genitalia present as the chlamydia infection, which may be asymptomatic or may resemble a gonorrhea infection.[11] Both are common causes of multiple other conditions including pelvic inflammatory disease and urethritis.[5] C. trachomatis is the single most important infectious agent associated with blindness (trachoma), and it also affects the eyes in the form of inclusion conjunctivitis and is responsible for about 19% of adult cases of conjunctivitis.[6] C. trachomatis in the lungs presents as the chlamydia pneumoniae respiratory infection and can affect all ages.[21] PathogenesisElementary bodies are generally present in the semen of infected men and vaginal secretions of infected women.[18] When they come into contact with a new host cell, the elementary bodies bind to the cell via interaction between adhesins on their surface and several host receptor proteins and heparan sulfate proteoglycans.[3] Once attached, the bacteria inject various effector proteins into the host cell using a type three secretion system.[3] These effectors trigger the host cell to take up the elementary bodies and prevent the cell from triggering apoptosis.[3] Within 6 to 8 hours after infection, the elementary bodies transition to reticulate bodies and a number of new effectors are synthesized.[3] These effectors include a number of proteins that modify the inclusion membrane (Inc proteins), as well as proteins that redirect host vesicles to the inclusion.[3] 8 to 16 hours after infection, another set of effectors are synthesized, driving acquisition of nutrients from the host cell.[3] At this stage, the reticulate bodies begin to divide, coinciding with the expansion of the inclusion.[3] If several elementary bodies have infected a single cell, their inclusions will fuse at this point to create a single large inclusion in the host cell.[3] From 24 to 72 hours after infection, reticulate bodies transition to elementary bodies which are released either by lysis of the host cell or extrusion of the entire inclusion into the host genital tract.[3] Virulence FactorsThe chlamydial plasmid, a DNA molecule existing separately from the genome of C. trachomatis, functions to enhance genetic diversity via the genes encoded.[22]The plasmid gene protein 3 (pgp3) has been linked to the establishment of persistent infection within the genital tract by suppressing the host immune response.[23] Polymorphic outer membrane proteins (Pmp proteins) on the surface of C. trachomatis use tropism to bind specific host cell receptors, which in turn initiates infection.[24] Pmp proteins B, D, and H have been most associated with eliciting a pro-inflammatory response through the release of cytokines.[25] CPAF (Chlamydia Protease-like Activity Factor) functions by preventing the host from triggering the proper immune response. C. trachomatis use of CPAF targets and cleaves proteins that restructure the Golgi apparatus and activate DNA repair so that C. trachomatis is able to use the host cell machinery and proteins to its advantage.[26] PresentationMost people infected with C. trachomatis are asymptomatic. However, the bacteria can present in one of three ways: genitourinary (genitals), pulmonary (lungs), and ocular (eyes).[7] Genitourinary cases can include genital discharge, vaginal bleeding, itchiness (pruritus), painful urination (dysuria), among other symptoms.[8] Often, symptoms are similar to those of a urinary tract infection.[citation needed] When C. trachomatis presents in the eye in the form of trachoma, it begins by gradually thickening the eyelids and eventually begins to pull the eyelashes into the eyelid.[27] In the form of inclusion conjunctivitis, the infection presents with redness, swelling, mucopurulent discharge from the eye, and most other symptoms associated with adult conjunctivitis.[6] C. trachomatis may latently infect the chorionic villi tissues of pregnant women, thereby impacting pregnancy outcome.[28] PrevalenceThree times as many women are diagnosed with genitourinary C. trachomatis infections as men. Women aged 15–19 have the highest prevalence, followed by women aged 20–24, although the rate of increase of diagnosis is greater for men than for women. Risk factors for genitourinary infections include unprotected sex with multiple partners, lack of condom use, and low socioeconomic status living in urban areas.[7] Pulmonary infections can occur in infants born to women with active chlamydia infections, although the rate of infection is less than 10%.[8] Ocular infections take the form of inclusion conjunctivitis or trachoma, both in adults and children. About 84 million worldwide develop C. trachomatis eye infections and 8 million are blinded as a result of the infection.[9] Trachoma is the primary source of infectious blindness in some parts of rural Africa and Asia and is a neglected tropical disease that has been targeted by the World Health Organization for elimination by 2020.[29] Inclusion conjunctivitis from C. trachomatis is responsible for about 19% of adult cases of conjunctivitis.[6] TreatmentTreatment depends on the infection site, age of the patient, and whether another infection is present. Having a C. trachomatis and one or more other sexually transmitted infections at the same time is possible. Treatment is often done with both partners simultaneously to prevent reinfection. C. trachomatis may be treated with several antibiotic medications, including azithromycin, erythromycin, ofloxacin,[11] and tetracycline. Tetracycline is the most preferred antibiotic to treat C.trachomatis and has the highest success rate. Azithromycin and doxycycline have equal efficacy to treat C. trachomatis with 97 and 98 percent success, respectively. Azithromycin is dosed as a 1 gram tablet that is taken by mouth as a single dose, primarily to help with concerns of non-adherence.[12] Treatment with generic doxycycline 100 mg twice a day for 7 days has equal success with expensive delayed-release doxycycline 200 mg once a day for 7 days.[12] Erythromycin is less preferred as it may cause gastrointestinal side effects, which can lead to non-adherence. Levofloxacin and ofloxacin are generally no better than azithromycin or doxycycline and are more expensive.[12] If treatment is necessary during pregnancy, levofloxacin, ofloxacin, tetracycline, and doxycycline are not prescribed. In the case of a patient who is pregnant, the medications typically prescribed are azithromycin, amoxicillin, and erythromycin. Azithromycin is the recommended medication and is taken as a 1 gram tablet taken by mouth as a single dose.[12] Despite amoxicillin having fewer side effects than the other medications for treating antenatal C. trachomatis infection, there have been concerns that pregnant women who take penicillin-class antibiotics can develop a chronic persistent chlamydia infection.[30] Tetracycline is not used because some children and even adults can not withstand the drug, causing harm to the mother and fetus.[12] Retesting during pregnancy can be performed three weeks after treatment. If the risk of reinfection is high, screening can be repeated throughout pregnancy.[11] If the infection has progressed, ascending the reproductive tract and pelvic inflammatory disease develops, damage to the fallopian tubes may have already occurred. In most cases, the C. trachomatis infection is then treated on an outpatient basis with azithromycin or doxycycline. Treating the mother of an infant with C. trachomatis of the eye, which can evolve into pneumonia, is recommended.[11] The recommended treatment consists of oral erythromycin base or ethylsuccinate 50 mg/kg/day divided into four doses daily for two weeks while monitoring for symptoms of infantile hypertrophic pyloric stenosis (IHPS) in infants less than 6 weeks old.[12] There have been a few reported cases of C.trachomatis strains that were resistant to multiple antibiotic treatments. However, as of 2018, this is not a major cause of concern as antibiotic resistance is rare in C.trachomatis compared to other infectious bacteria.[31] Laboratory testsChlamydia species are readily identified and distinguished from other Chlamydia species using DNA-based tests. Tests for Chlamydia can be ordered from a doctor, a lab or online.[32] Most strains of C. trachomatis are recognized by monoclonal antibodies (mAbs) to epitopes in the VS4 region of MOMP.[33] However, these mAbs may also cross-react with two other Chlamydia species, C. suis and C. muridarum.[citation needed]
ResearchStudies have revealed antibiotic resistance in Chlamydia trachomatis. Mutations in the 23S rRNA gene, including A2057G and A2059G, have been identified as significant contributors to resistance against azithromycin, a commonly used treatment. This resistance is linked to treatment failures and persistent infections, necessitating ongoing research into alternative antibiotics, such as moxifloxacin, as well as non-antibiotic approaches like bacteriophage therapy. These innovations aim to combat resistance while reducing the overall burden of antibiotic misuse, which has been closely associated with the rise of resistant strains in C. trachomatis populations.[37] Additionally, diagnostic improvements have played a vital role in identifying C. trachomatis infections more efficiently. Nucleic acid amplification tests (NAATs), such as DNA- and RNA-based tests, have shown high sensitivity and specificity, making them the gold standard for detecting asymptomatic infections. NAATs have facilitated broader screening programs, particularly in high-risk populations, and are integral to public health initiatives aimed at controlling the spread of C. trachomatis. Research continues into point-of-care diagnostic tools, which promise faster results and greater accessibility, especially in low-resource settings.[38] In the area of vaccine development, creating an effective vaccine for C. trachomatis has proven challenging due to the complex immune responses the bacterium elicits. Subunit vaccines, which target outer membrane proteins like MOMP (Major Outer Membrane Protein) and polymorphic membrane proteins (Pmp), are being explored in both animal models and early human trials. While these vaccines show promise in inducing partial immunity in murine models, further research is needed to evaluate their efficacy in humans. The goal is to develop a vaccine that can prevent reinfection without causing harmful inflammatory responses.[39] HistoryC. trachomatis was first described in 1907 by Stanislaus von Prowazek and Ludwig Halberstädter in scrapings from trachoma cases.[40][17] Thinking they had discovered a "mantled protozoan", they named the organism "Chlamydozoa" from the Greek "Chlamys" meaning mantle.[17] Over the next several decades, "Chlamydozoa" was thought to be a virus as it was small enough to pass through bacterial filters and unable to grow on known laboratory media.[17] However, in 1966, electron microscopy studies showed C. trachomatis to be a bacterium.[17] This is essentially due to the fact that they were found to possess DNA, RNA, and ribosomes like other bacteria. It was originally believed that Chlamydia lacked peptidoglycan because researchers were unable to detect muramic acid in cell extracts.[41] Subsequent studies determined that C. trachomatis synthesizes both muramic acid and peptidoglycan, but relegates it to the microbe's division septum and does not utilize it for construction of a cell wall.[42][43] The bacterium is still classified as gram-negative.[44] C. trachomatis agent was first cultured and isolated in the yolk sacs of eggs by Tang Fei-fan et al. in 1957.[45] This was a significant milestone because it became possible to preserve these agents, which could then be used for future genomic and phylogenetic studies. The isolation of C. trachomatis coined the term isolate to describe how C. trachomatis has been isolated from an in vivo setting into a "strain" in cell culture.[46] Only a few "isolates" have been studied in detail, limiting the information that can be found on the evolutionary history of C. trachomatis.[45][47] EvolutionIn the 1990s it was shown that there are several species of Chlamydia. Chlamydia trachomatis was first described in historical records in Ebers papyrus written between 1553 and 1550 BC.[48] In the ancient world, it was known as the blinding disease trachoma. The disease may have been closely linked with humans and likely predated civilization.[49] It is now known that C. trachomatis comprises 19 serovars which are identified by monoclonal antibodies that react to epitopes on the major outer-membrane protein (MOMP).[50] Comparison of amino acid sequences reveals that MOMP contains four variable segments: S1,2 ,3 and 4. Different variants of the gene that encodes for MOMP, differentiate the genotypes of the different serovars. The antigenic relatedness of the serovars reflects the homology levels of DNA between MOMP genes, especially within these segments.[51] Furthermore, there have been over 220 Chlamydia vaccine trials done on mice and other non-human host species to target C. muridarum and C. trachomatis strains. However, it has been difficult to translate these results to the human species due to physiological and anatomical differences. Future trials are working with closely related species to humans.[52] See alsoReferences
Further reading
External links |
||||||||||||||||||||||||||||||||||||||||||||||||